The New Era of Visual Proteomics: Mapping the Cell in 3D
For decades, structural biology was like trying to understand a city by looking at a few isolated buildings. We could see a single protein in high resolution, but we had no idea how it behaved in the chaotic, crowded environment of a living cell. That is changing rapidly.
The shift toward cryo-electron tomography (cryo-ET) is turning the tide. Instead of isolating proteins in a test tube, researchers are now capturing “snapshots” of the cellular interior in its native state. The trend is moving toward what experts call Visual Proteomics—the ability to identify and map every single protein complex within a cell’s 3D architecture.
Imagine a Google Maps for the cell. We aren’t just seeing where the proteins are; we are seeing how they touch, bend, and interact in real-time. This level of detail is critical for understanding neurodegenerative diseases, where protein misfolding creates “traffic jams” in the neurons.
Mechanobiology: Engineering the Cell’s Internal Architecture
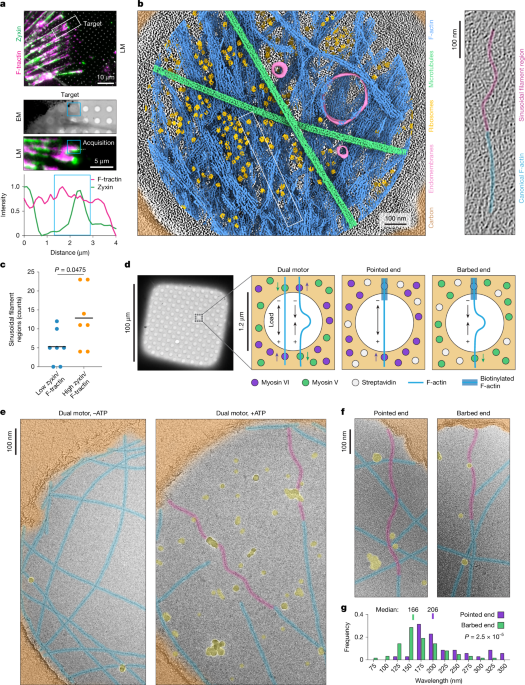
We used to think of the cytoskeleton—the network of actin and myosin—as a passive scaffold. We now know We see a highly dynamic machine that senses and responds to physical force. The discovery of F-actin supercoiling under mechanical strain reveals that the cell doesn’t just “feel” force; it stores it like a spring.
The future of medicine lies in manipulating these mechanical switches. In cancer research, for instance, the “stiffness” of a tumor’s environment often dictates how aggressively the cancer spreads. By understanding the molecular mechanics of how proteins like $alpha$-catenin sense tension, we can develop drugs that “trick” a cell into thinking it is in a stable environment, potentially halting metastasis.
Beyond pathology, this is the frontier of bio-hybrid robotics. By controlling the force-generation of myosin motors, engineers are beginning to create synthetic tissues that can contract and move with the precision of biological muscle, paving the way for advanced prosthetics.
AI-Powered Microscopy: Beyond the Human Eye
The biggest bottleneck in structural biology hasn’t been the microscopes—it’s been the data. A single cryo-ET session can produce terabytes of noisy, grainy images that would capture a human years to analyze.
The integration of Deep Learning, specifically Denoising Autoencoders (DAE) and U-Net architectures, is automating the “picking” and segmentation of particles. We are moving toward a world of Autonomous Microscopy, where the AI identifies the most interesting structures in a sample and directs the microscope to zoom in on them in real-time.
This isn’t just about speed; it’s about accuracy. AI can now distinguish between a genuine protein complex and a piece of carbon contamination with a precision that exceeds human capability. As these models evolve, we will see “predictive imaging,” where AI suggests the likely structure of a protein before the image is even fully processed.
The Rise of the “Digital Twin” for Proteins
Parallel to imaging is the rise of coarse-grained molecular dynamics. By creating digital twins of filaments and motors, scientists can simulate how a protein will react to a specific drug or a physical force without ever touching a pipette.
This “in silico” approach reduces the cost of drug discovery by orders of magnitude. Instead of testing 10,000 compounds in a lab, researchers can simulate them on a digital actin filament and only test the five most promising candidates.
Frequently Asked Questions
What is Cryo-ET, and why is it better than traditional EM?
Traditional electron microscopy often requires chemicals that distort the cell. Cryo-ET flash-freezes the sample in liquid ethane, preserving the cell in a “vitrified” state that looks exactly like it did while alive.

How does AI help in studying proteins?
AI removes “noise” from images (denoising) and automatically identifies protein shapes (segmentation), allowing scientists to analyze thousands of molecules instead of just a few.
What is the significance of actin supercoiling?
It shows that the cell’s skeleton can change its physical shape to respond to force, which acts as a mechanical signal to regulate cell growth, movement, and repair.
Join the Conversation
Do you think AI will eventually replace the need for physical lab experiments in structural biology? Or will the “wet lab” always be the gold standard?
Share your thoughts in the comments below or subscribe to our newsletter for weekly insights into the future of biotech!